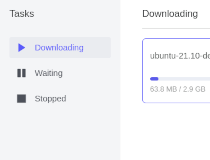
Protein shapes can be investigated experimentally, using x-ray crystallography, but this is an expensive, error-prone and time-consuming laboratory operation, so in recent years researchers have been pursuing entirely computer-based solutions. Sequencing proteins is now fairly routine, but the key to biology is the three-dimensional shape of the protein - how a protein “folds”. Each protein is specified as a string of amino acids (A, C, T or G), typically several thousand long. Examples in human biology include actin and myosin, the proteins that enable muscles to work, and hemoglobin, the basis of red blood that carries oxygen to cells. Model of human nuclear pore complex, built using AlphaFold2 credit: Agnieszka Obarska-Kosinska, NatureProteins are the workhorses of biology. After 40 days of training by playing games with itself, AlphaGo Zero was as far ahead of Ke Jie as Ke Jie is ahead of a good amateur player. Later that same year, a new DeepMind program named “AlphaGo Zero” defeated the earlier Alpha Go program 100 games to zero. Then in May 2017, a computer program named “AlphaGo,” developed by DeepMind, defeated Ke Jie, a 19-year-old Chinese Go master thought to be the world’s best human Go player. Many observers did not expect Go-playing computer programs to beat the best human players for many years, if ever. The ancient Chinese game of Go is notoriously complicated, with strategies that can only be described in vague, subjective terms. 
First, we will briefly review some of DeepMind’s earlier achievements, upon which their new matrix work is based. Recent DeepMind achievementsĪs mentioned above, researchers at DeepMind, a research subsidiary of Alphabet (Google’s parent), have devised a machine learning-based search program that has not only reproduced many of the specific results in the literature, but has also discovered a few schemes, for certain specific size classes, that are even more efficient than the best known methods. By decomposing each of these matrices into half-sized (i.e., $n \times n$) submatrices $A_)$, compared with $O(n^3)$ for the conventional scheme. Consider matrices $A, B$ and $C$, which, for simplicity in the presentation here, may each be assumed to be of size $2n \times 2n$ for some integer $n$ (although the algorithm is also valid for more general size combinations). Strassen’s algorithm for matrix multiplication 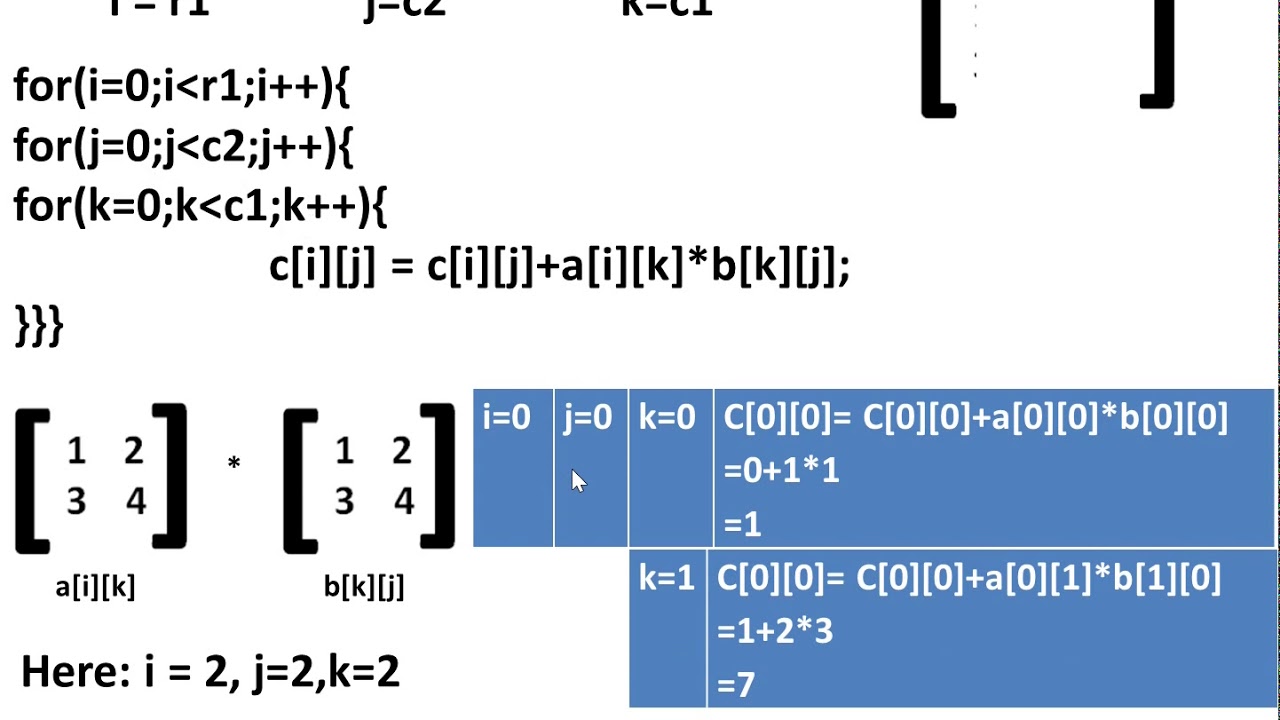
In this article, we present an introduction to these fast matrix multiplication methods, and then describe the latest results. The latest development here is that researchers at DeepMind, a research subsidiary of Alphabet (Google’s parent), have devised a machine learning-based program that has not only reproduced many of the specific results in the literature, but has also discovered a few schemes, for certain specific size classes, that are even more efficient than the best known methods. 
For a good overview of these methods, together with some numerical analysis and other considerations, see Higham2022. In the years since Strassen’s discovery, numerous other researchers have found even better schemes for certain specific matrix size classes. German mathematician Volker Strassen was the first to show this, in 1969, by exhibiting a scheme that yields practical speedups even for moderately sized matrices. What could be more simple and straightforward? Thus it may come as a surprise to some the basic scheme is not the most efficient. Most of us learn the basic scheme for matrix multiplication in high school.
0 Comments
Leave a Reply. |

 RSS Feed
RSS Feed
